Biomolecular Function Design
Associate Professor
- OKUDA, Aya, e-mail: okuda.aya.5r<atmark>kyoto-u.ac.jp
Assistant Professor
- KUROKAWA, Minami e-mail: kurokawa.minami.2x<atmark>kyoto-u.ac.jp
* replace <atmark> with @rri.kyoto-u.ac.jp
Our laboratory aims to elucidate the molecular mechanisms underlying diverse biological phenomena through the analysis of protein structure and function. We also develop a wide range of applications by modifying and regulating protein function.
1) Functional and Structural Analysis of Oxidative Protein Folding Enzymes
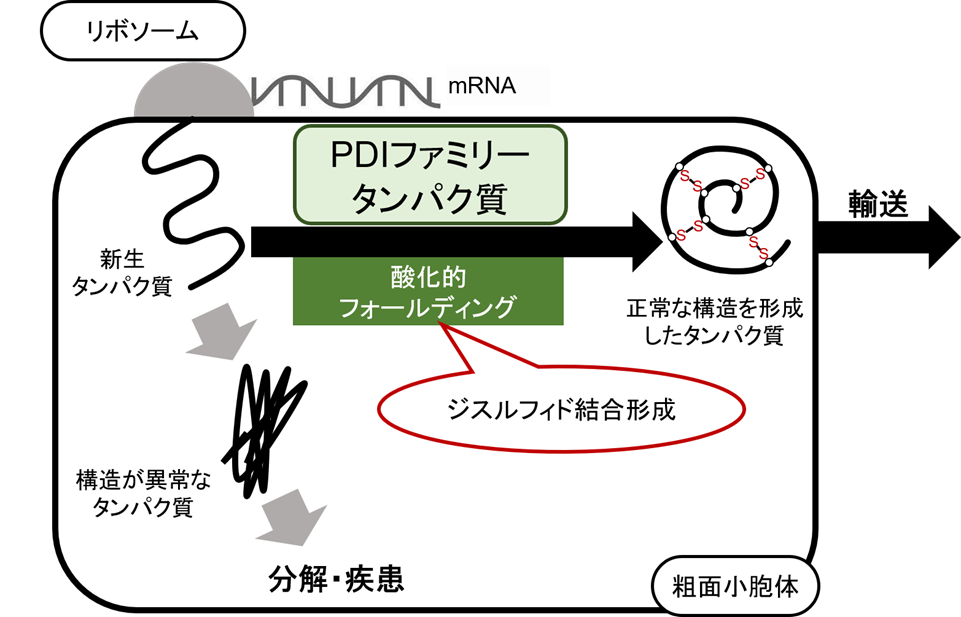
Secretory proteins, which play a crucial role in maintaining vital functions in eukaryotes, are synthesized on ribosomes bound to the rough endoplasmic reticulum (ER). Ribosomes read the genetic information encoded in mRNA and translocate nascent polypeptides into the lumen of the ER. Once properly folded into its native structure, it is transported to various sites where it performs its function. However, proteins that fail to form the correct three-dimensional structure are either degraded by the endoplasmic reticulum’s quality control mechanisms or accumulate, potentially causing neurodegenerative diseases. Therefore, proper protein folding is essential for correct transport and function, and elucidating the mechanisms of protein folding in the endoplasmic reticulum remains an important challenge.
Much of the three-dimensional structure formation of proteins synthesized in the endoplasmic reticulum involves the formation of disulfide bonds. This is referred to as oxidative folding, and this process is carried out by a group of oxidative folding enzymes known as the protein disulfide isomerase (PDI) family. In our laboratory, we aim to elucidate the mechanism of oxidative folding by analyzing the functions and structures of human and soybean PDI family proteins through biochemical and structural analysis using X-ray and neutron solution scattering.
2) Development of Protein Ligation Technique
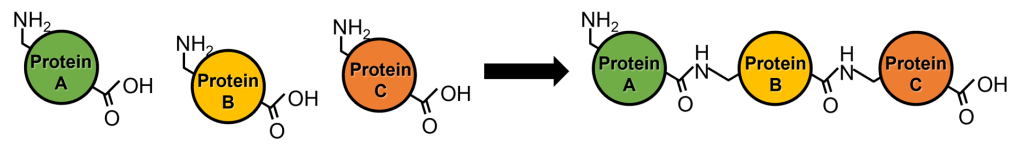
Protein ligation refers to the technique of covalently connecting two or more polypeptides or proteins via peptide bonds or similar mechanisms. Typically, during ribosomal protein synthesis, amino acids are sequentially linked via peptide bonds starting from the N-terminus to synthesize the full-length protein. In contrast, protein ligation involves connecting proteins that have already been synthesized.
This technique enables the incorporation of various biochemical and biophysical probes, such as fluorescent probes and non-natural amino acids, into proteins, in addition to structural analysis using NMR and small-angle neutron scattering (SANS) focused on specific amino acid sequences or domains through stable isotope labeling. Furthermore, this technique enables advances in peptide and protein engineering, such as the cyclization of peptides and proteins and the production of proteins that would otherwise be toxic in their full-length form.
However, significant challenges remain in achieving efficie