Biomolecular Structure
Professor
- INOUE Rintaro, e-mail: inoue.rintaro.5w*
Associate Professor
- CHATAKE Toshiyuki, e-mail: chatake.toshiyuki.6x*
Assistant Professors
- KITA Akiko, e-mail: kita.akiko.4u*
- KAWAGUCHI Akio, e-mail: kawaguchi.akio.6s*
◆replace* with @kyoto-u.ac. jp
Biomacromolecules function as nanoscale machines that constitute living systems, and their diverse structures and intrinsic fluctuations (dynamics) are closely linked to their functions. In our laboratory, we employ a complementary combination of X-ray and neutron scattering, spectroscopic techniques, and computational analyses to achieve precise characterization of the structures and dynamics of biomacromolecules.
1. Visualization of domain motions in multi-domain proteins through integrated analysis of neutron scattering and computational methods
It is well recognized that domain motions in multi-domain proteins are closely linked to their functions; however, methods to directly analyze such motions have long remained undeveloped. By exploiting the fact that the energy of slow neutrons is comparable to the thermal fluctuations of proteins, we successfully measured domain motions in multi-domain proteins using neutron scattering. Furthermore, by integrating computational analyses, we achieved visualization of these domain motions and demonstrated that they correspond to essential dynamic modes directly involved in protein function.
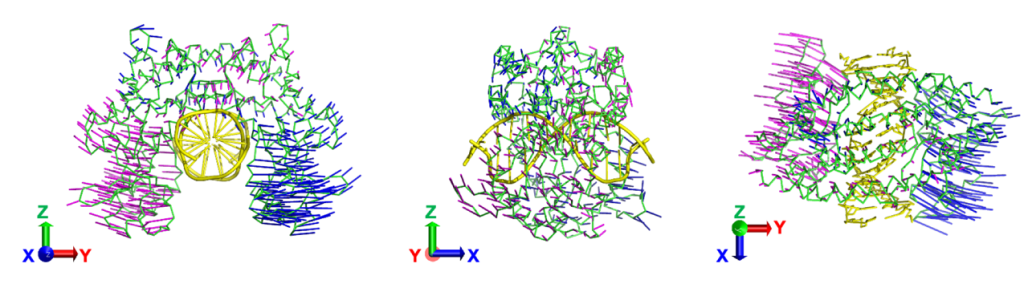
Fig. 1 Domain motions visualized through integrated analysis of neutron scattering and computational methods.
2. D/H contrast neutron protein crystallography
The neutron scattering lengths of hydrogen and deuterium are different (bH = 3.8 fm, bD = 6.6 fm). The difference between the neutron Fourier maps of H2O-solvent and D2O-solvent protein crystals provides density contrasts of D/H exchangeable atoms (D/H contrast) of amino and hydroxyl groups in protein and water. D/H contrast neutron protein crystallography is expected to improve the efficiency and accuracy of hydrogen analysis. In our laboratory, the algorithm for calculating the D/H contrast in real space has been developed, and D/H neutron crystallographic analyses of proteins have been carried out.
Fig. 2. Neutron D/H contrast map in the solvent region of a protein crystal. (a) Two types of water molecules with different dynamic behavior. (b) Neutron contrast map superimposed on conventional neutron map.
3. Structural study of novel and/or biologically interested protein
To know the protein structures including H atoms by neutron diffraction method as described, it is necessary to determine their three-dimensional structures with no H atoms. The structure analyses of novel and/or biologically interesting proteins are in progress with collaborators, utilizing X-ray diffraction method that has no upper limits of the molecular weight.